Large-scale structural variation (SV) is pervasive in the human genome, both in healthy individuals and in tumor cells. Numerous methods have been developed to detect such variants, most of which rely on the information provided by molecularly paired reads. Even the most sophisticated methods, however, still generate numerous false positives. A new study in Nature Genetics describes an innovative, population-based method to improve the accuracy of SV calling. In their introduction, Handsaker et al offer four main causes underlying false positives in SV calls:
- Sequencing errors, which occur more frequently in next-generation sequencing data and exhibit both random and platform-specific bias distributions.
- Chimeric molecules, in which read pairs linking two non-continguous segments of DNA masquerade as SVs. Sequencing libraries can contain millions of such fragments, which represent ~1% of sequence reads.
- Read depth variation, which fluctuates across the genome for both biological and technical reasons.
- Genome repeats, which confound most short-read aligners even when read pairing information is available.
These issues are exacerbated in population-scale sequencing, which often yields lower coverage across large numbers of samples. As more genomes are sequenced, false positives accumulate faster than real variants do. However, the authors hypothesized that population-scale sequencing might enable new analytical approaches. Here, they describe three strategies to do just that: allele sharing, population heterogeneity, and allelic substitution.
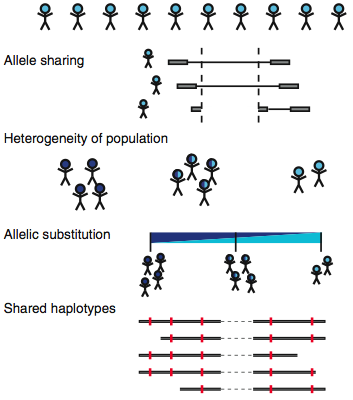
Credit: Nat. Genet. 768 (Handsaker et al, 2011)
Coherence Around Shared Alleles
Most of the variation in any given genome is shared, at some level, with other members of the population. Pilots from the 1,000 Genomes Project have shown that variants with appreciable allele frequencies (>1%) in the population will generally be shared by multiple samples if the pool is sufficiently sized. Further, for medical sequencing projects, causative variants should be enriched among cases even if they’re rare at the population level. The authors sought to exploit shared variation wherever it could be found, without filtering out singleton variants. In essence, they looked for evidence of similar deletion alleles (measured by larger-than-expected insert sizes for read pairs) across multiple samples. The idea was that random chimeric events should be specific to a single library, whereas SVs reflecting true variation should persist across multiple libraries from multiple samples. Looking in the 1,000 Genomes data, the authors found that 89% of the SVs had evidence across multiple genomes.
However, it became clear that allele coherence by itself was an insufficient criterion for SV calling, because even after it was applied, there were ten times the number of expected SVs according to extrapolations of copy number data.
Heterogeneity in Populations
Next, the authors sought to use allele heterogeneity in their sample populations to distinguish real variants, which should be present in some individuals, but not others. For each deletion, they performed a chi squared test of the number of read pairs supporting or not supporting the variant across 168 genomes. The resulting p-value, or heterogeneity statistic, was consistently low for “control” deletions that were known to be real by copy number data. Many of the loci that had passed the shared allele coherence test, but failed the heterogeneity statistic, were flanked by homologous sequences that caused aligners to mis-place reads; copy number data suggested that few such cases represented real variants.
Copy Number Correlations
To bolster the support for putative SVs, the authors evaluated the relationship between predicted deletions and copy number depth for the reference allele. In theory, if the variants represented true deletions, there should be a corresponding drop in coverage. In many cases, there was no such correlation; further review showed that many of these loci bore cryptic polymorphisms (often small indels) that caused reads to mis-align to nearby, paralogous sequences. Another cause of predictions that passed shared-allele and heterogeneity tests but failed the read depth correlation was transposon insertion polymorphisms not contained in the reference sequence. Reads from such insertions often mapped to nearby paralogous sequences, thereby falsely supporting large deletions of the intervening sequences.
Breakpoint Resolution and Genotype Determination
By combining data across all individuals found to have a structural allele in common, it was possible to localize the breakopints of deletions with resolutions of 1-20 bp. Many types of information in population-scale sequencing data – paired-end alignments, read depth, and breakpoint-spanning reads – can supply partial information about the genotype state of SVs in individuals. The authors developed a Bayesian framework to integrate this information into an integrated measurement of relative likelihood that the sequence data from each genome arose from each potential SV allele at that locus. Comparisons of these inferred genotypes to copy number data and (where available) high-resolution array genotype data supported a high accuracy for the method, and showed that the confidence score tracked with accuracy.
For deletions smaller than 300 bp, few genotypes could be inferred with high confidence. To resolve these, the authors utilized haplotypes formed by SNPs and SVs together. Most common SVs characterized to date have been shown to segregate with common SNP haplotypes; by employing imputation algorithms and haplotype information, the authors were able to extend this approach to resolve many low-confidence genotypes. The resulting calls were consistent with the features of sequence data and also fit the haplotype structure of the population. The authors genotyped 13,826 of the deletion polymorphisms identified by the 1,000 Genomes project (ranging in size from 48 bp to 960 kbp), with an average call rate of 94.1%. This was ten times as many deletions as could be genotyped by combination SNP-CNV arrays that were designed for genome-wide association studies.
In summary, Handsaker and colleagues have presented strategies that could help develop new analytical approaches as sequencing is extended to large populations. Together with SNP- and small indel-detection algorithms, these approaches will help realize the full potential of population-scale sequencing.
References
Handsaker RE, Korn JM, Nemesh J, & McCarroll SA (2011). Discovery and genotyping of genome structural polymorphism by sequencing on a population scale. Nature genetics, 43 (3), 269-76 PMID: 21317889
Dan, I remain amazed at how you churn through these dense papers. Bravo! Do these strategies from population genomics help us with SV detection in cancer genomes in any way?
That would be nice, wouldn’t it? Unfortunately, most of these strategies rely on principles of inherited variation – e.g., “polymorphic” SVs that are segregating in the population. In tumors, acquired SVs don’t have to occur at sites of known polymorphisms – they likely arise completely sporadically. Across a population of tumors of the same type, however, we might be able to apply some of these principles to identify recurrent events (even if the alleles are different).